
ERC DINAMIX – Real-time diffusion NMR analysis of mixtures
DINAMIX is a project led by Jean-Nicolas Dumez aiming at developing NMR methods for the analysis of out-of-equilibrium mixtures. DINAMIX has been funded by a 1.5 M€ ERC Starting Grant (n° 801774, 2019-2024).
5-year project
2019 - 2024
€ 1 499 337
ERC funds
228
person-months effort over the 5 years
8
internal team members
In order to describe, understand, and eventually control chemical reactions, scientists need to know what is happening in reaction mixtures.
Nuclear magnetic resonance (NMR) is the chemist’s most powerful tool to "see" which molecules are in a sample. However, the available options are typically a comprehensive picture on a completed reaction, and partial snapshots throughout the reaction. Using concepts derived from magnetic resonance imaging (MRI) and recent instrumentation, we are developing methods to provide a more complete picture of solution mixtures that evolve in time.
In more details
Chemical samples often come as solution mixtures. While advanced analytical methods exist for samples at equilibrium, the information on components and their interactions that may be accessed for out-of-equilibrium mixtures is much more limited. These include, for example, reaction mixtures in chemical synthesis and catalysis, as well as biochemical enzymatic reactions. The DINAMIX project aim at providing additional molecular-level information on out-of-equilibrium mixtures.
The project relies on diffusion nuclear magnetic resonance (NMR) spectroscopy, a powerful method that separates the spectra of mixtures’ components and identifies interactions, in correlation with structural insight provided by NMR observables. A spatial parallelisation is used to approach that makes it possible to acquire data in less than a second, instead of several minutes with conventional methods. This also has the potential to boost the power of hyperpolarisation methods such as dissolution dynamic nuclear polarisation (D-DNP) for mixtures analysis.
D-DNP indeed provides NMR sensitivity enhancements of up to 4 orders of magnitude, which however last only for a short time in solution. The methods developed will be used for applications in reaction monitoring in catalytic organic synthesis and biochemistry.
Project team
Past members
Hélène BONIN
Project manager
Finances, human ressources, management and reporting
(2019-2022)
33.Clean 1H NMR Spectra of Products Directly from Batch and Flow Reaction Mixtures
Authors: Yuliia Horbenko, Martin Jaudronnet, Nour El Sabbagh, Margherita Bazzoni, Aurélie Bernard, Mathias Nilsson, Patrick Giraudeau, François-Xavier Felpin, and Jean-Nicolas Dumez
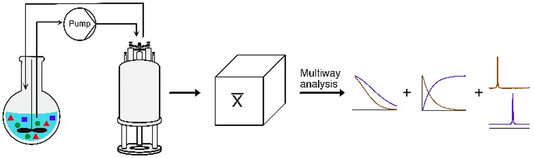
32.Fast Multidimensional Flow Nuclear Magnetic Resonance at High Field for Real-Time Reaction Monitoring and Flow Synthesis
Authors: Margherita Bazzoni, Yuliia Horbenko, Nour El Sabbagh, Achille Marchand, Magdalena Grochowska-Tatarczak, Aurélie Bernard, Patrick Giraudeau, François-Xavier Felpin, and Jean-Nicolas Dumez
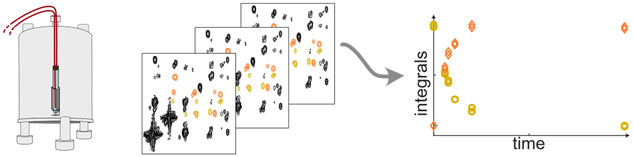
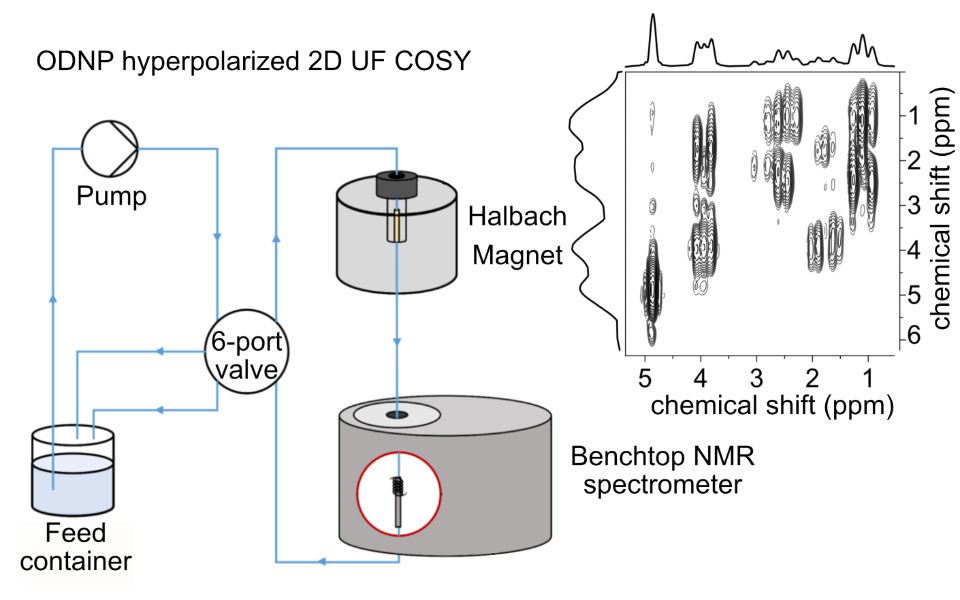
31.Ultrafast 2D benchtop NMR spectroscopy enhanced by flow Overhauser dynamic nuclear polarization
Authors: J. Mandral, J. Phuong, J. Farjon, P. Giraudeau, K. Münnemann, J.-N. Dumez

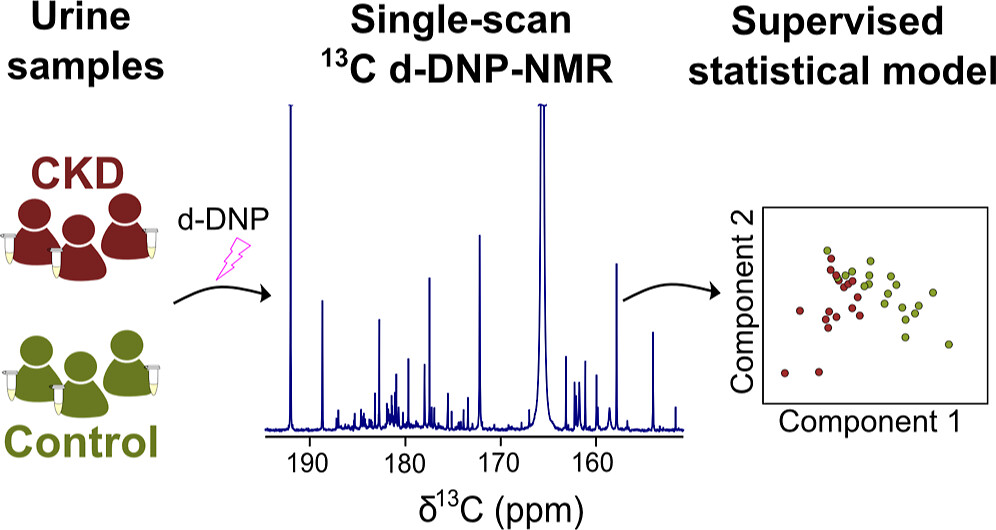
29. Hyperpolarized 13C NMR Metabolomics of Urine Samples at Natural Abundance Applied to Chronic Kidney Disease, doi: 10.1021/jacs.4c12607
Authors: V. Ribay, B. Charrier, M. Croyal, B. Cariou S. Hadjadj, J. Boccard, C. Cannet, J.-N. Dumez, M.P.M. Letertre, P. Giraudeau
28. Pure Shift NMR in Continuous Flow, doi: 10.1002/chem.202403385
Authors: M. Bazzoni, A. Régheasse, E. Caytan, F.-X. Felpin, P. Giraudeau, A. Bernard, R. W. Adams, G. A. Morris, M. Nilsson, J.-N. Dumez
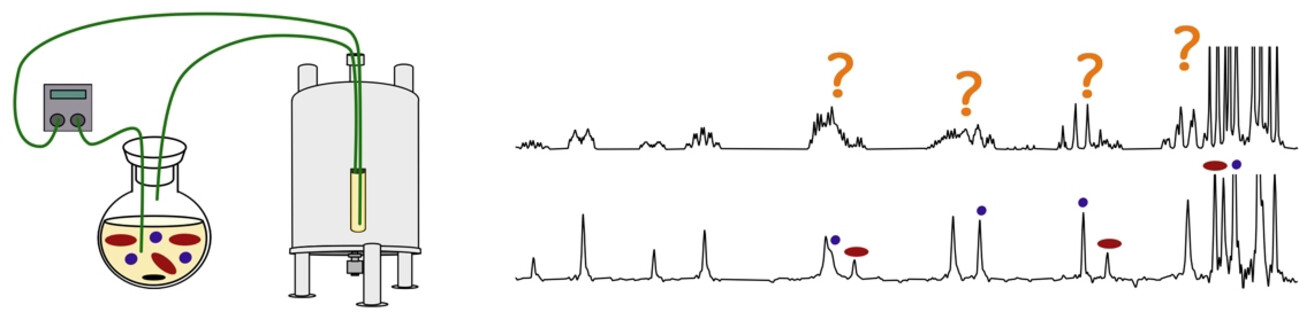
27. Autonomous reaction self-optimization using in-line high-field NMR spectroscopy (link)
Authors: N. El Sabbagh, M. Bazzoni, Y. Horbenko, D. Cortes-Borda, A. Bernard, P. Giraudeau, F.-X. Felpin and J.-N. Dumez
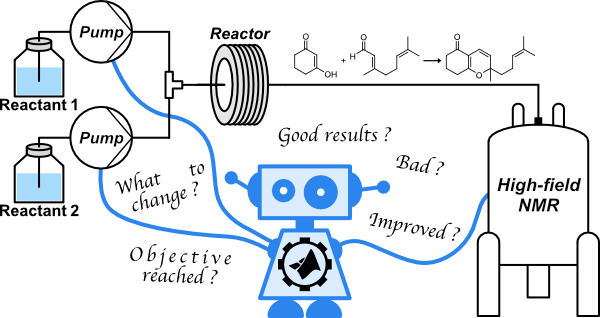
This work shows how in-line detection by high-field NMR combined with a feedback look can be used to drive the autonomous self optimisation of reaction conditions.
26. Spatially encoded pure-shift diffusion-ordered NMR spectroscopy yielded by chirp excitation (https://doi.org/10.1016/j.jmr.2024.107628)
Authors: Rituraj Mishra, Jonathan R.J. Yong, Corentin Jacquemmoz, Benjamin Lorandel, Mohammadali Foroozandeh, Jean-Nicolas Dumez
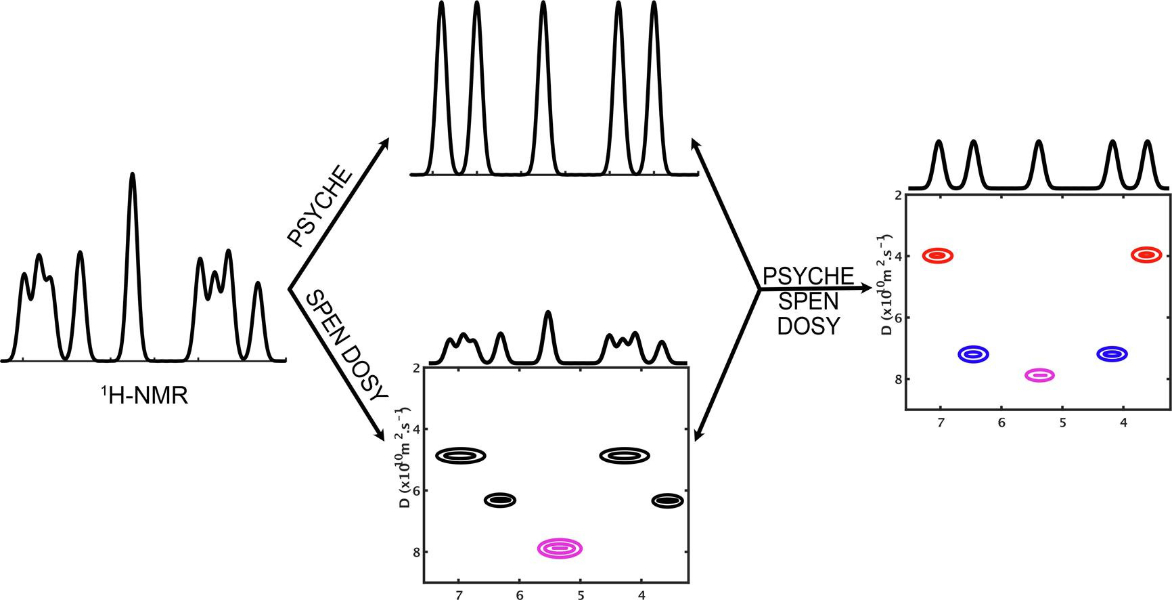
We show that the built-in position-dependent diffusion attenuation of PSYCHE NMR can be used to accelerate the acquisition of pure-shift DOSY data, using spatially resolved acquisition.
25. Optimization of heteronuclear ultrafast 2D NMR for the study of complex mixtures hyperpolarized by dynamic nuclear polarization (link)
Authors: Clément Praud, Victor Ribay, Arnab Dey, Benoît Charrier, Joris Mandral, Jonathan Farjon, Jean-Nicolas Dumez, and Patrick Giraudeau
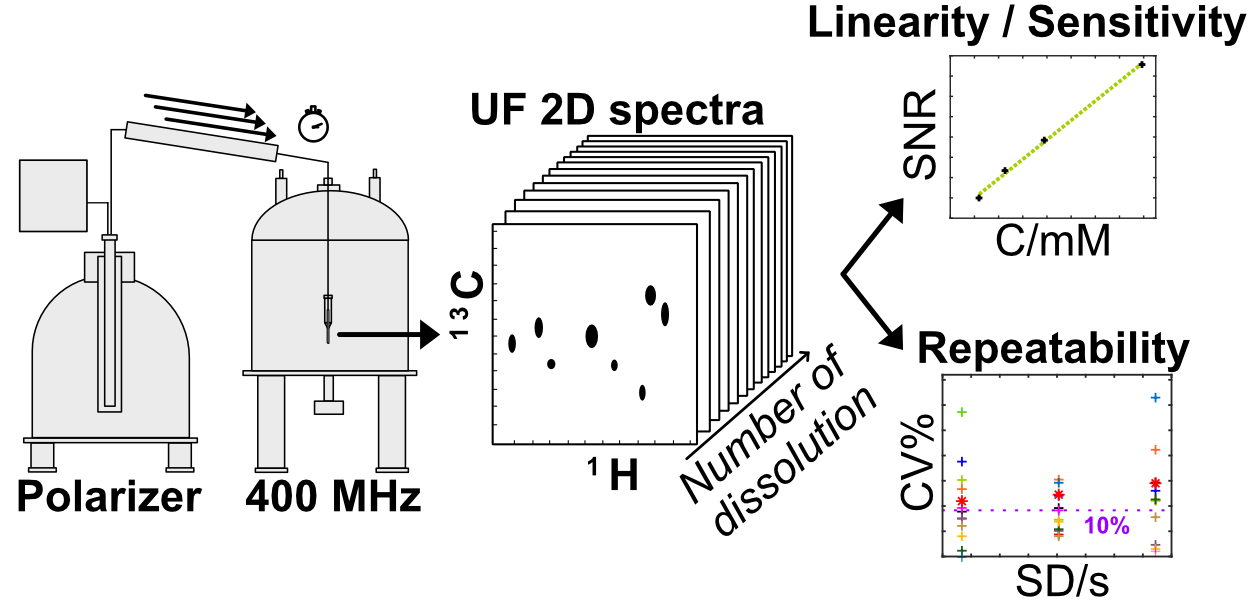
This work provides a rigorous characterisation of the sensitivity and repeatability of the combination of dissolution dynamic nuclear polarisation and ultrafast 2D NMR, which are important for analytical applications.
24. Hyperpolarized 1H and 13C NMR Spectroscopy in a Single Experiment for Metabolomics (link)
Authors: Arnab Dey, Benoît Charrier, Victor Ribay, Jean-Nicolas Dumez and Patrick Giraudeau
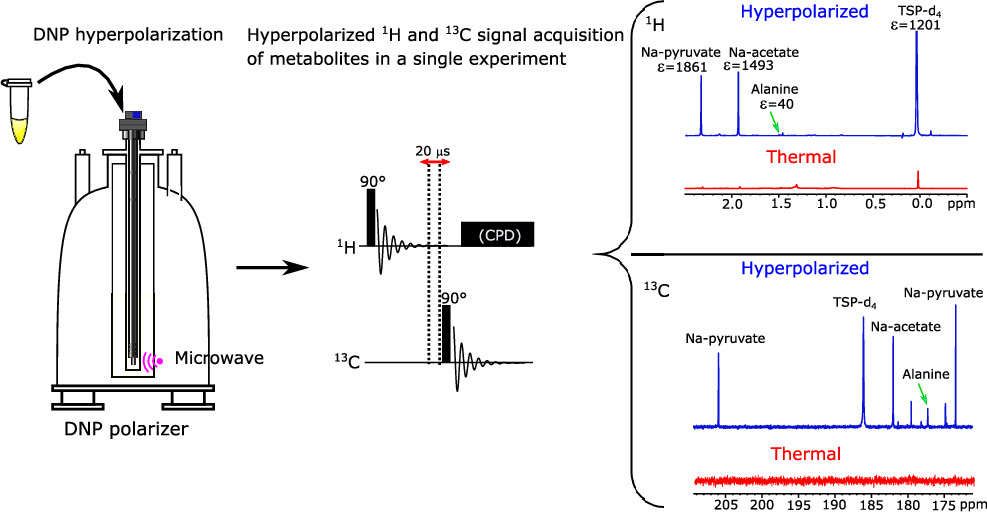
This work shows that hyperpolarised 13C and 1H 1D NMR spectra can be acquired in a single dissolution dynamic nuclear polarisation experiment, with signal enhancements of several orders of magnitude.
23. Single-Scan Ultraselective NMR Experiments with Preserved Sensitivity (link)
Authors: Margherita Bazzoni, Rituraj Mishra, Jean-Nicolas Dumez
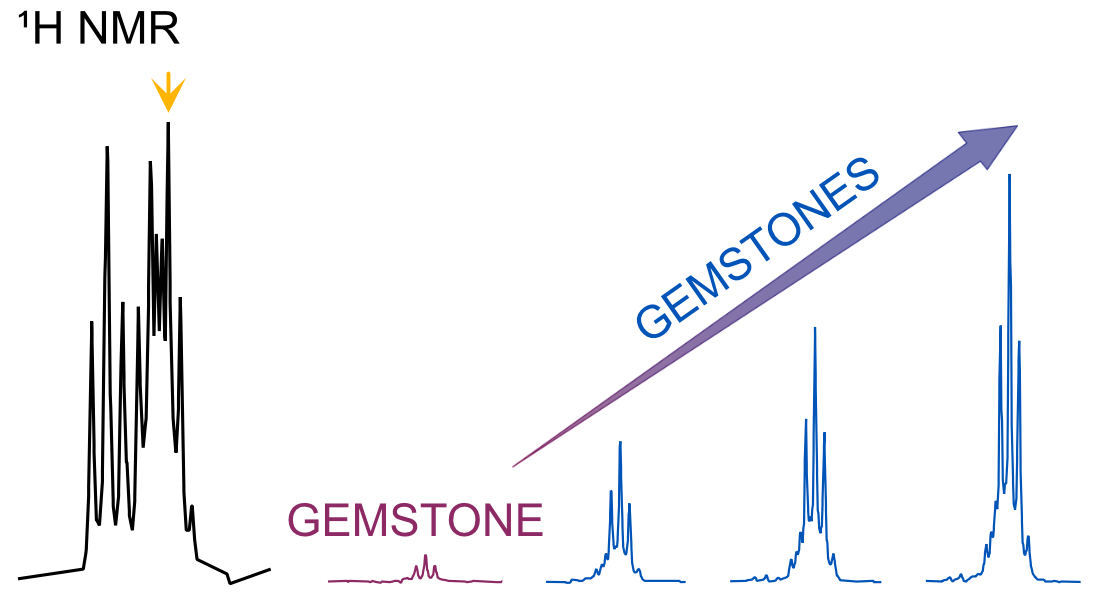
This work show that the sensitivity of fast ultraselective experiments, which address a single multiplet even when it overlap with other multiplets, can be improved significantly in cases where molecular translational diffusion has significant effects.
22. Solution-state 2D NMR spectroscopy of mixtures hyperpolarized using optically polarized crystals (link)
Authors: Anna J. Parker, Arnab Dey, Mohammed Usman Qureshi, Jakob M. Steiner, John W. Blanchard, Jochen Scheuer, Nikolas Tomek, Stefan Knecht, Felix Josten, Christoph Muller, Patrick Hautle, Ilai Schwartz, Patrick Giraudeau, Tim Eichhorn, Jean-Nicolas Dumez
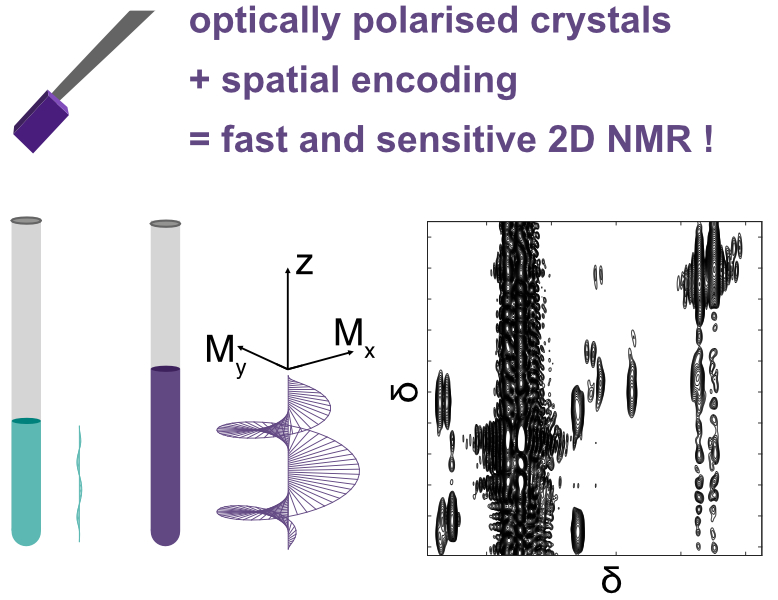
In this work, we show that 2D correlation spectra can be acquired in a single-scan, from a mixture of substrates that are hyperpolarised using optically polarised crystals.
21. Broadband ultrafast 2D NMR spectroscopy for online monitoring in continuous flow (link)
Authors: Célia Lhoste, Margherita Bazzoni, Justine Bonnet, Aurélie Bernard, François-Xavier Felpin, Patrick Giraudeau, Jean-Nicolas Dumez
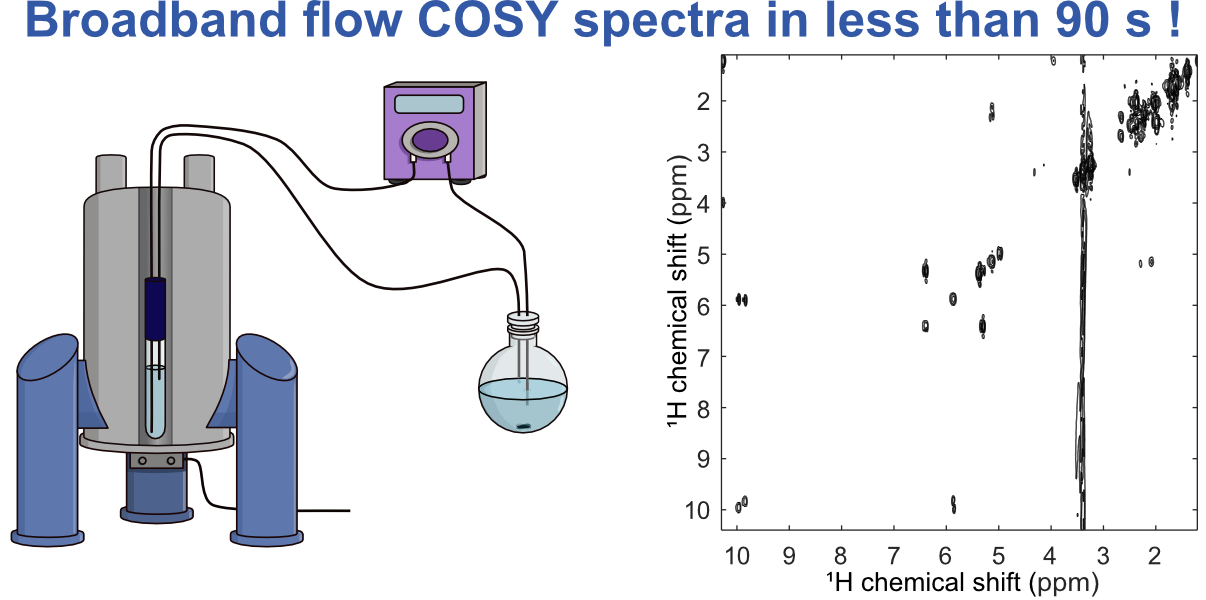
We show that broadband NMR spectra, covering the full 1H spectral width, can be acquired in just a few scans for a flowing sample, using interleaved ultrafast NMR acquisition
20. Accounting for gradient non-uniformity in spatially-encoded diffusion-ordered NMR spectroscopy (link)
Authors: Benjamin Lorandel, Rituraj Mishra, Oksana Cazimajou, Achille Marchand, Aurélie Bernard, Jean-Nicolas Dumez
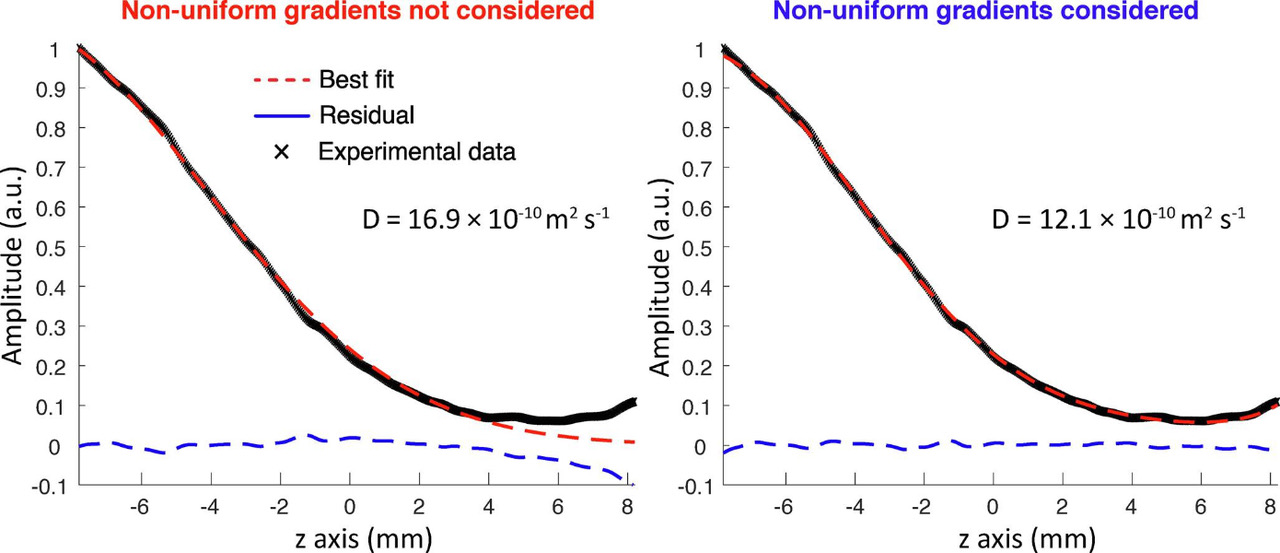
This work explains how to account for gradient non uniformity in the processing of SPEN DOSY data, and show the resulting improvement in accuracy
19. Book chapter: Fast 2D NMR for reaction and process monitoring (link)
Authors: Margherita Bazzoni, Benjamin Lorandel, Célia Lhoste, Patrick Giraudeau, Jean-Nicolas Dumez
18. Book chapter: Fast 2D NMR for metabolomics (link)
Authors: Clément Praud, Marine P.M. Letertre, Arnab Dey, Jean-Nicolas Dumez, Patrick Giraudeau
17. Hyperpolarized 13C NMR Spectroscopy of Urine Samples at Natural Abundance by Quantitative Dissolution Dynamic Nuclear Polarization (link)
Authors: Victor Ribay, Arnab Dey, Benoît Charrier, Clément Praud, Joris Mandral, Jean-Nicolas Dumez, Marine P.M. Letertre, and Patrick Giraudeau
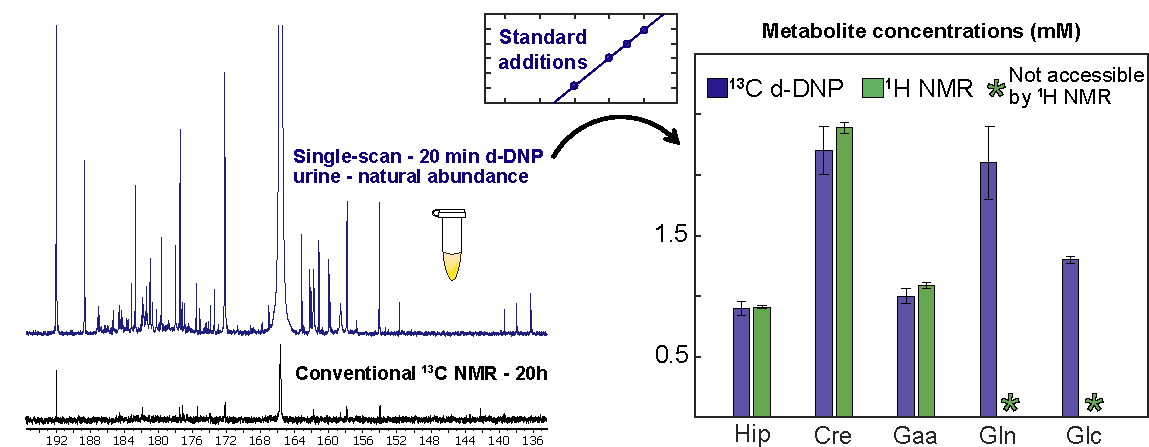
This article reports the first d-DNP-enhanced 13C NMR analysis of a biofluid -urine- at natural abundance, offering unprecedented resolution and sensitivity for this challenging type of sample.
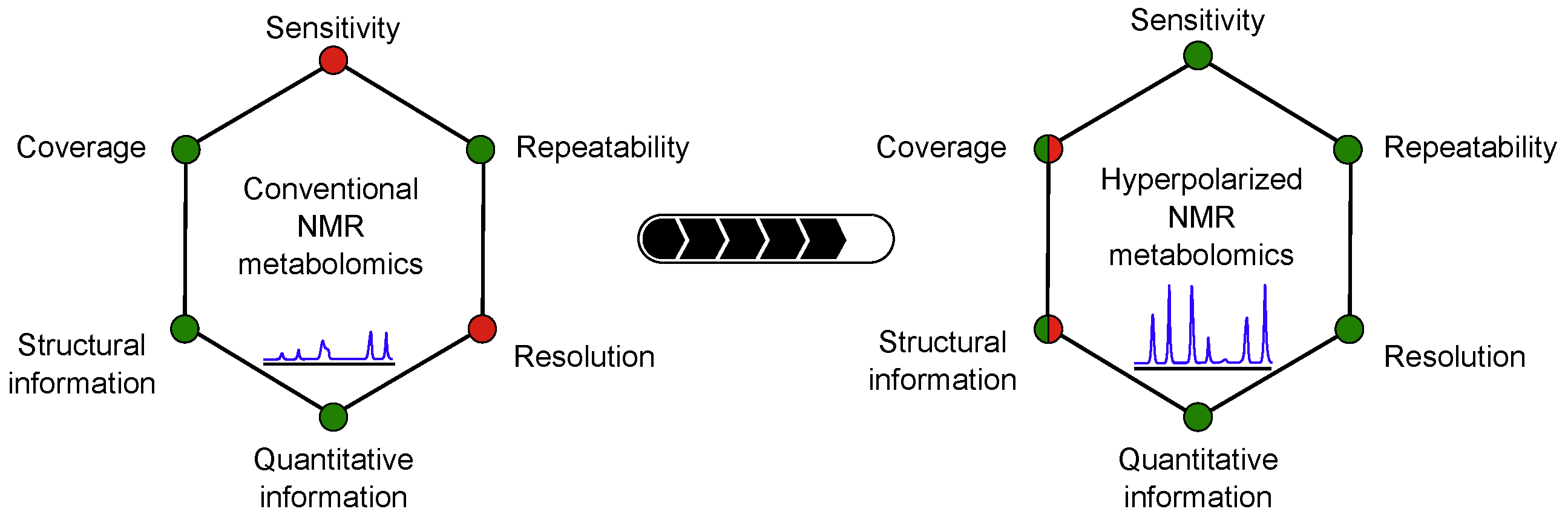
16. Hyperpolarized NMR metabolomics (link)
Authors: Victor Ribay, Clément Praud, Marine P. M. Letertre, Jean-Nicolas Dumez and Patrick Giraudeau
15. In-line Multidimensional NMR Monitoring of Photochemical Flow Reactions (Link)
Authors: Margherita Bazzoni, Célia Lhoste, Justine Bonnet, Kouakou Eric Konan, Aurélie Bernard, Patrick Giraudeau, François-Xavier Felpin, Jean-Nicolas Dumez
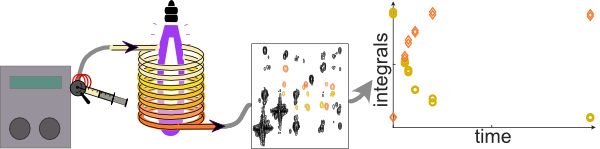
This work demonstrates the in-line monitoring of a flow photochemical reaction using 1D and ultrafast 2D NMR methods at high magnetic field.
14. Theoretical analysis of flow effects in spatially encoded diffusion NMR (Link)
Authors: Rituraj Mishra, Jean-Nicolas Dumez
This work presents a detailed theoretical and numerical analysis of flow effects in spatially encoded diffusion NMR spectroscopy.
13. NMR methods for the analysis of mixtures (link)
Author: Jean-Nicolas Dumez
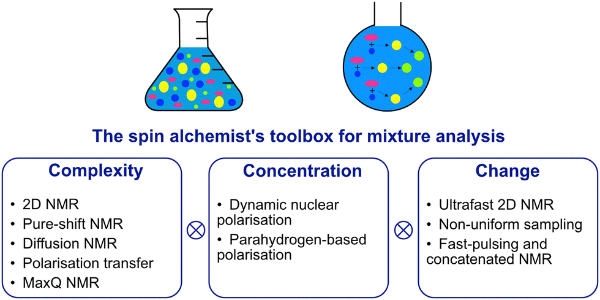
A feature article that describes NMR methods to tackle the complexity, low concentration and changing nature of mixtures.
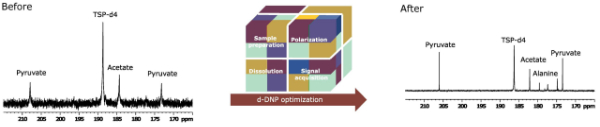
12. Fine optimization of a dissolution dynamic nuclear polarization experimental setting for 13C NMR of metabolic samples (link)
Authors: Arnab Dey, Benoît Charrier, Karine Lemaitre, Victor Ribay, Dmitry Eshchenko, Marc Schnell, Roberto Melzi, Quentin Stern, Samuel F. Cousin, James G. Kempf, Sami Jannin, Jean-Nicolas Dumez, and Patrick Giraudeau
11. Online Reaction Monitoring with Fast and Flow-Compatible Diffusion NMR Spectroscopy (link)
Authors: Achille Marchand, Rituraj Mishra, Aurélie Bernard, Jean-Nicolas Dumez
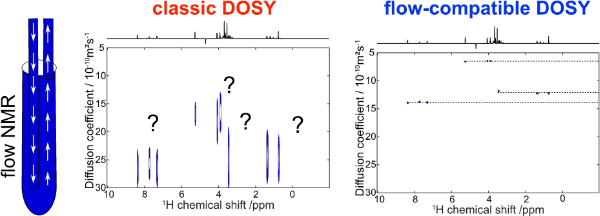
We describe a fast and flow-compatible diffusion NMR experiment and use it to monitor an organic chemical reaction.
10. Ultrafast 2D NMR for the analysis of complex mixtures (link)
Authors: Célia Lhoste, Benjamin Lorandel, Clément Praud, Achille Marchand, Rituraj Mishra, Arnab Dey, Aurélie Bernard, Jean-Nicolas Dumez, Patrick Giraudeau
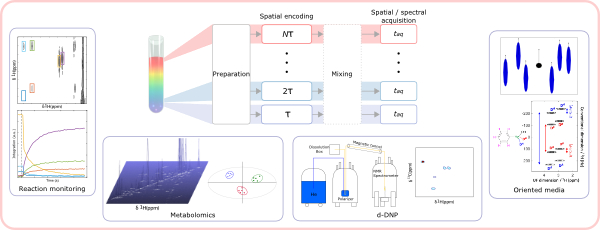
This review article describes ultrafast 2D NMR and its applications in the fields of monitoring, metabolomics, hyperpolarization and oriented media.
9. Quadratic spacing of the effective gradient area for spatially encoded diffusion NMR (link)
Authors: Rituraj Mishra and Jean-Nicolas Dumez
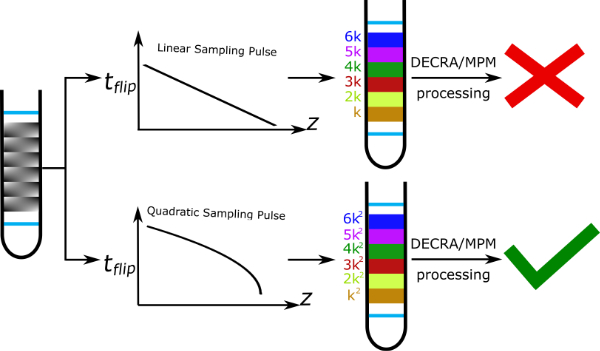
We illustrate the mechanism of a frequency-swept pulse that achieves quadratic spacing of the effective gradient area in ultrafast DOSY NMR, and show that makes it possible to use fast processing methods.
8. Ultrafast 2D 1H-1H NMR spectroscopy of DNP-hyperpolarised substrates for the analysis of mixtures (link)
Authors: Kawarpal Singh, Corentin Jacquemmoz, Patrick Giraudeau, Lucio Frydman, Jean-Nicolas Dumez
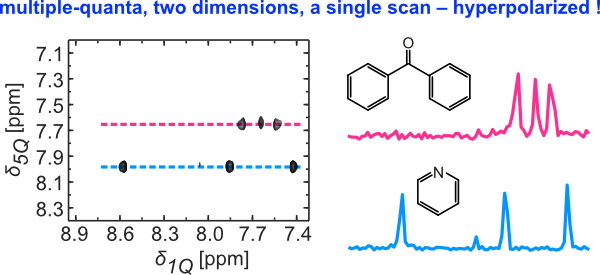
We show that TOCSY and multiple-quantum (MQ) 2D NMR spectra can be obtained for mixtures of substrates hyperpolarised by dissolution dynamic nuclear polarisation (D-DNP).
7. Optimisation of spatially encoded diffusion ordered NMR spectroscopy for the analysis of mixtures (link)
Authors: Corentin Jacquemmoz, Rituraj Mishra, Ludmilla Guduff, Carine van Heijenoort, Jean-Nicolas Dumez
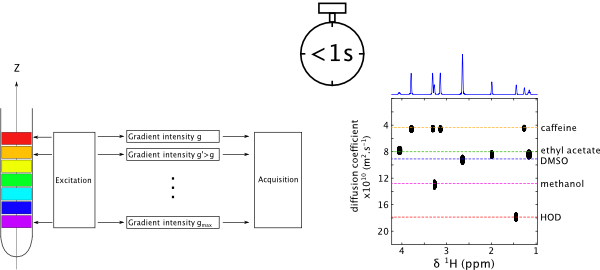
This article presents the unusual principles and implementation of the SPEN DOSY experiment, which is particularly useful for systems that evolve in time.
6. Ultrafast diffusion-based unmixing of 1H NMR spectra (link)
Authors: Rituraj Mishra, Achille Marchand, Corentin Jacquemmoz, and Jean-Nicolas Dumez
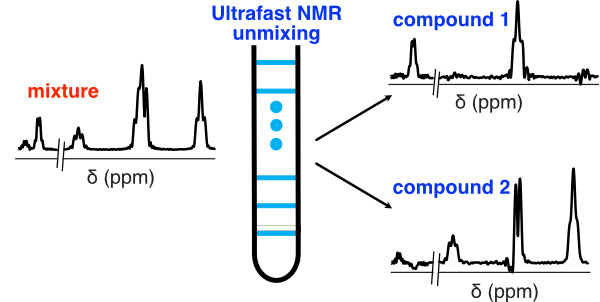
We show that the NMR spectra of components in a mixture can be separated using 2D data acquired in less than one second, and an algorithm that is executed in just a few seconds.
5. Online reaction monitoring by single-scan 2D NMR under flow conditions (link)
Authors: Corentin JACQUEMMOZ, François GIRAUD, Jean-Nicolas DUMEZ.
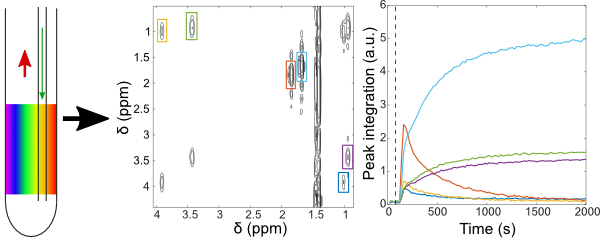
Here we show that time series of single-scan ultrafast 2D NMR (UF2DNMR) spectra can be collected to monitor solution mixtures that circulate in a flow unit at high field.
4. Hyperpolarized NMR Metabolomics at Natural 13C Abundance (link)
Authors: Arnab Dey, Benoît Charrier, Estelle Martineau, Catherine Deborde, Elodie Gandriau, Annick Moing, Daniel Jacob, Dmitry Eshchenko, Marc Schnell, Roberto Melzi, Dennis Kurzbach, Morgan Ceillier, Quentin Chappuis, Samuel F. Cousin, James G. Kempf, Sami Jannin, Jean-Nicolas Dumez and Patrick Giraudeau
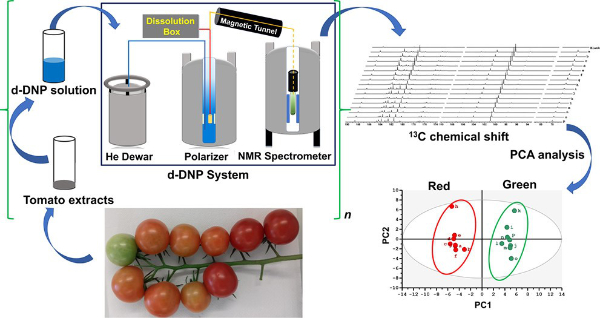
This article introduces a new approach, based on hyperpolarized 13C NMR at natural abundance, that circumvents the sensitivity drawback of 13C NMR for metabolomics.
3. Frequency-swept pulses for ultrafast spatially encoded NMR
Author: Jean-Nicolas Dumez
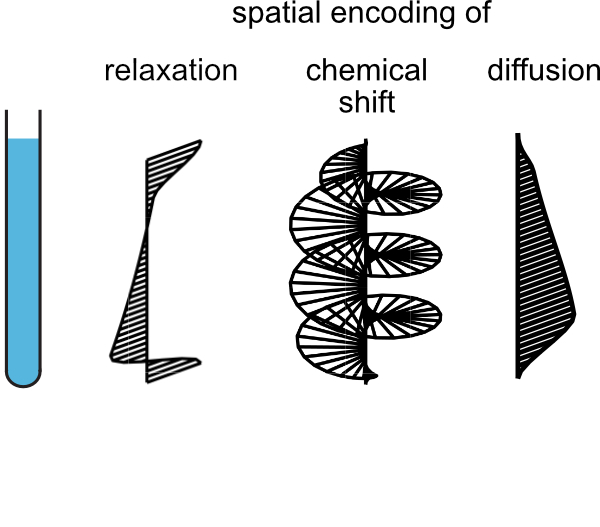
This article describes the spatial encoding of relaxation, chemical shift and diffusion in a common framework and discusses directions for future developments.
2. Application and methodology of dissolution dynamic nuclear polarization in physical, chemical and biological contexts (link)
Authors: Sami Jannin, Jean-Nicolas Dumez, Patrick Giraudeau, Dennis Kurzbach
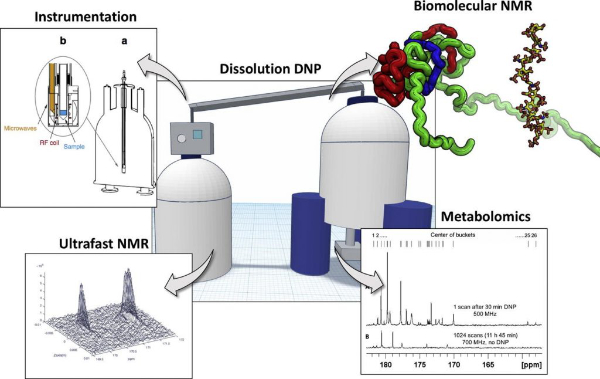
This perspectives article highlights possible avenues for developments and applications of d-DNP in biochemical and physicochemical studies.
1. Interleaved spatial/spectral encoding in ultrafast 2D NMR spectroscopy (link)
Authors: Bertrand Plainchont, Patrick Giraudeau, Jean-Nicolas Dumez
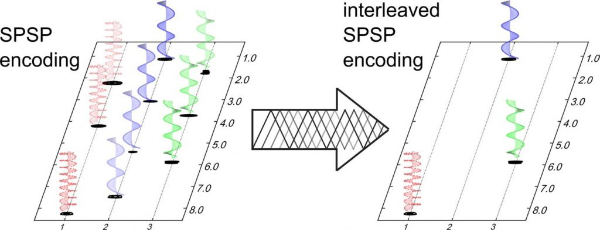
We analyze and further develop the strategy of using spatial/spectral pulses for spatial encoding, and introduce the concept of interleaving in excitation space, to address drawback of UF2DNMR.
